"""
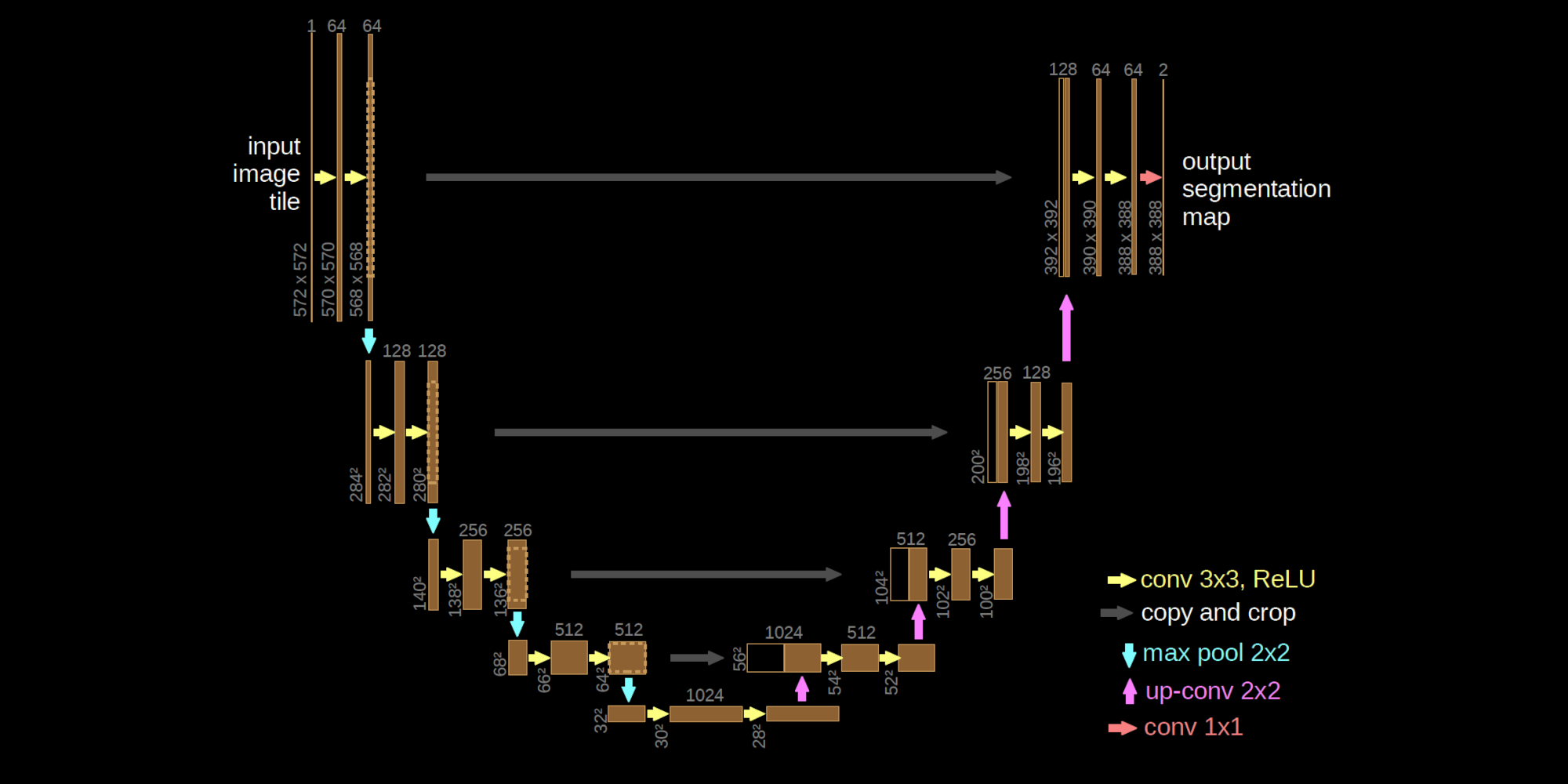
U-Net neural operator implemented with Equinox.
Architecture:
- Lifting DoubleConv
- Encoder: stride-2 Conv downsampling + DoubleConv at each level
- Decoder: ConvTranspose upsampling + skip-connection concat + DoubleConv
- 1x1 Conv projection to output channels
"""
import jax
import jax.numpy as jnp
import jax.tree_util as jtu
import equinox as eqx
from typing import Callable
class DoubleConv(eqx.Module):
conv_1: eqx.nn.Conv
conv_2: eqx.nn.Conv
activation: Callable
def __init__(
self,
num_spatial_dims: int,
in_channels: int,
out_channels: int,
activation: Callable,
*,
key,
):
c_1_key, c_2_key = jax.random.split(key)
self.conv_1 = eqx.nn.Conv(
num_spatial_dims,
in_channels,
out_channels,
kernel_size=3,
padding=1,
key=c_1_key,
)
self.conv_2 = eqx.nn.Conv(
num_spatial_dims,
out_channels,
out_channels,
kernel_size=3,
padding=1,
key=c_2_key,
)
self.activation = activation
def __call__(self, x: jax.Array) -> jax.Array:
x = self.activation(self.conv_1(x))
x = self.activation(self.conv_2(x))
return x
class UNet(eqx.Module):
lifting: DoubleConv
down_sampling_blocks: list[eqx.nn.Conv]
left_arc_blocks: list[DoubleConv]
right_arc_blocks: list[DoubleConv]
up_sampling_blocks: list[eqx.nn.ConvTranspose]
projection: eqx.nn.Conv
def __init__(
self,
num_spatial_dims: int,
in_channels: int,
out_channels: int,
hidden_channels: int,
num_levels: int,
activation: Callable,
*,
key,
):
key, lifting_key, projection_key = jax.random.split(key, 3)
self.lifting = DoubleConv(
num_spatial_dims,
in_channels,
hidden_channels,
activation,
key=lifting_key,
)
self.projection = eqx.nn.Conv(
num_spatial_dims,
hidden_channels,
out_channels,
kernel_size=1,
key=projection_key,
)
# Channel counts at each level: [C, 2C, 4C, ...]
channel_list = [hidden_channels * 2**i for i in range(num_levels + 1)]
self.down_sampling_blocks = []
self.left_arc_blocks = []
self.right_arc_blocks = []
self.up_sampling_blocks = []
for upper_ch, lower_ch in zip(channel_list[:-1], channel_list[1:]):
key, down_key, left_key, right_key, up_key = jax.random.split(key, 5)
self.down_sampling_blocks.append(
eqx.nn.Conv(
num_spatial_dims,
upper_ch,
upper_ch,
kernel_size=3,
stride=2,
padding=1,
key=down_key,
)
)
self.left_arc_blocks.append(
DoubleConv(num_spatial_dims, upper_ch, lower_ch, activation, key=left_key)
)
self.right_arc_blocks.append(
DoubleConv(num_spatial_dims, lower_ch, upper_ch, activation, key=right_key)
)
self.up_sampling_blocks.append(
eqx.nn.ConvTranspose(
num_spatial_dims,
lower_ch,
upper_ch,
kernel_size=3,
stride=2,
padding=1,
output_padding=1,
key=up_key,
)
)
def __call__(self, x: jax.Array) -> jax.Array:
x = self.lifting(x)
x_skips = []
# Encoder (left arc)
for down, left in zip(self.down_sampling_blocks, self.left_arc_blocks):
x_skips.append(x)
x = down(x)
x = left(x)
# Decoder (right arc)
for right, up in zip(
reversed(self.right_arc_blocks), reversed(self.up_sampling_blocks)
):
x = up(x)
# Equinox operates without a batch axis; channels are at axis 0
x = jnp.concatenate([x, x_skips.pop()], axis=0)
x = right(x)
return self.projection(x)
def count_parameters(model: eqx.Module) -> int:
"""Return the total number of trainable scalar parameters."""
return sum(p.size for p in jtu.tree_leaves(eqx.filter(model, eqx.is_array)))